Problem with ligand-RMSD (l-RMSD) calculation in the basic protein-protein docking tutorial - HADDOCK - BioExcel
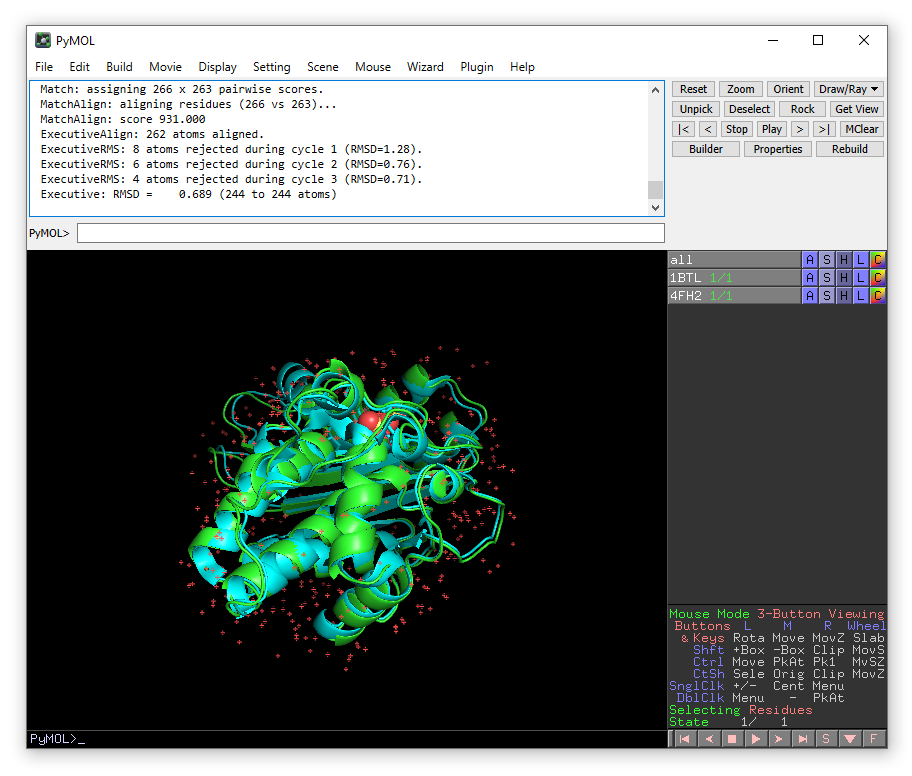
GitHub - tongalumina/rmsdca: PyMOL script to calculate backbone RMSD of two polypeptides of same origin

How to label the amino acids involved in the protein protein interaction using PyMol software, as shown in expected labels.png? | ResearchGate

How to calculate rmsd value of conformers by PyMol ( root mean square deviation calculation) - YouTube