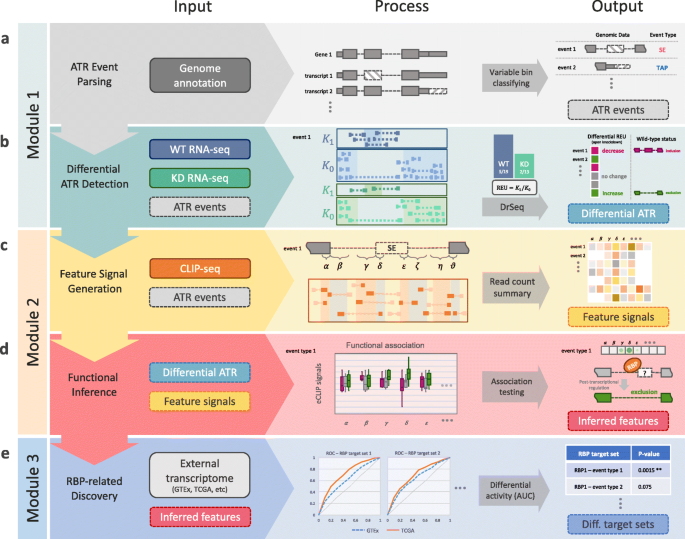
SURF: integrative analysis of a compendium of RNA-seq and CLIP-seq datasets highlights complex governing of alternative transcriptional regulation by RNA-binding proteins | Genome Biology | Full Text

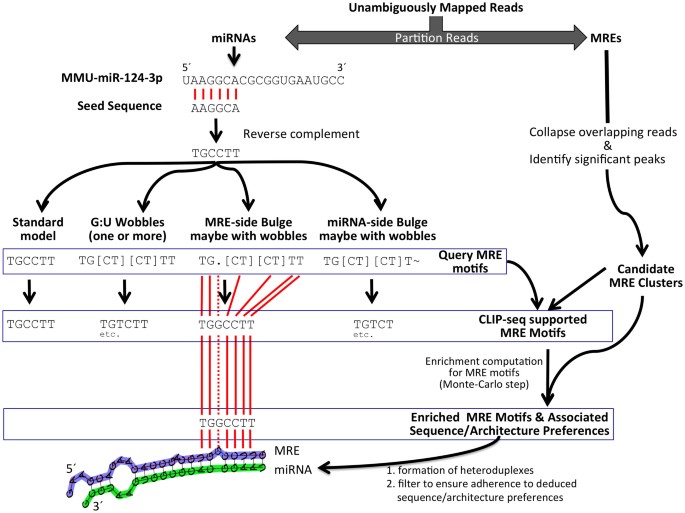
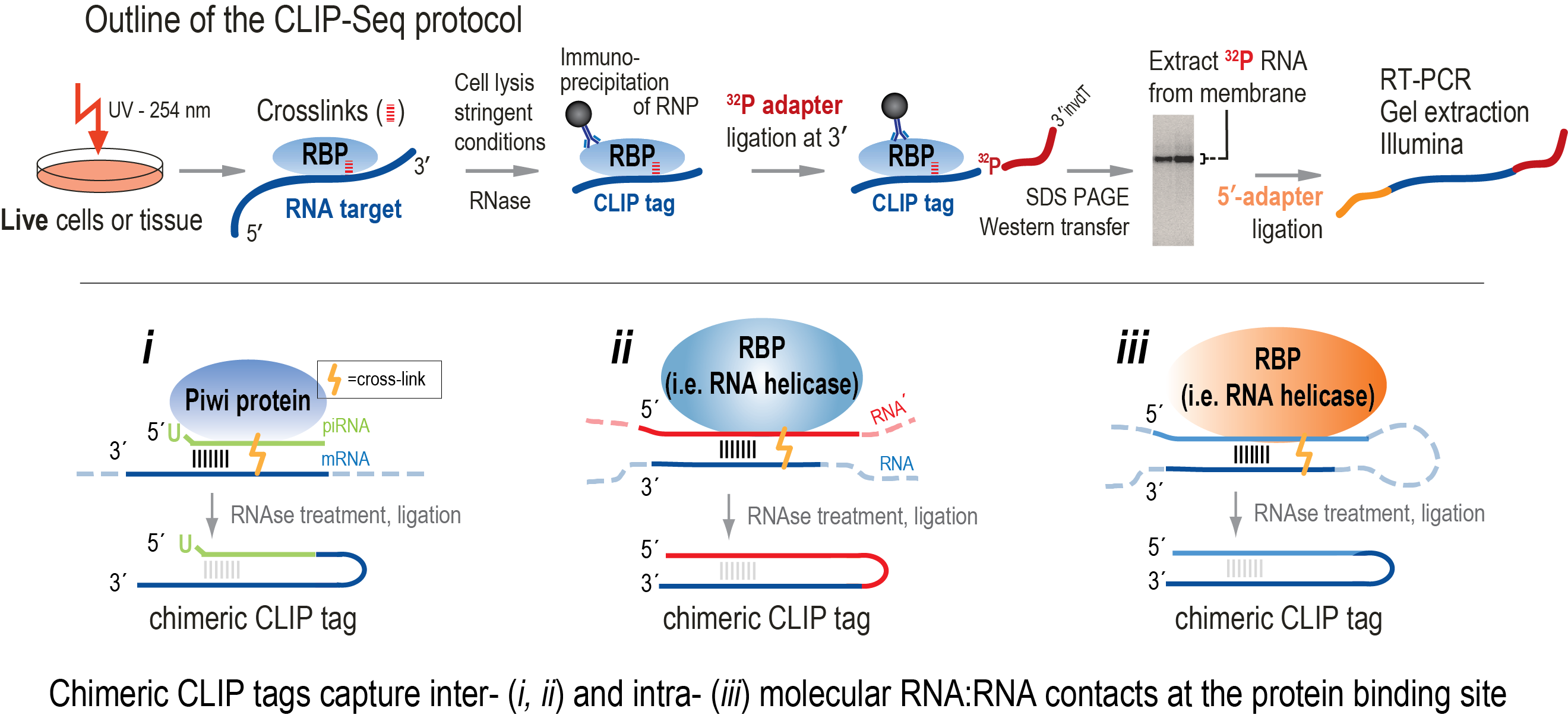
Pipeline for extracting MIWI CLIP-Seq features (see Section 2). The RNA... | Download Scientific Diagram
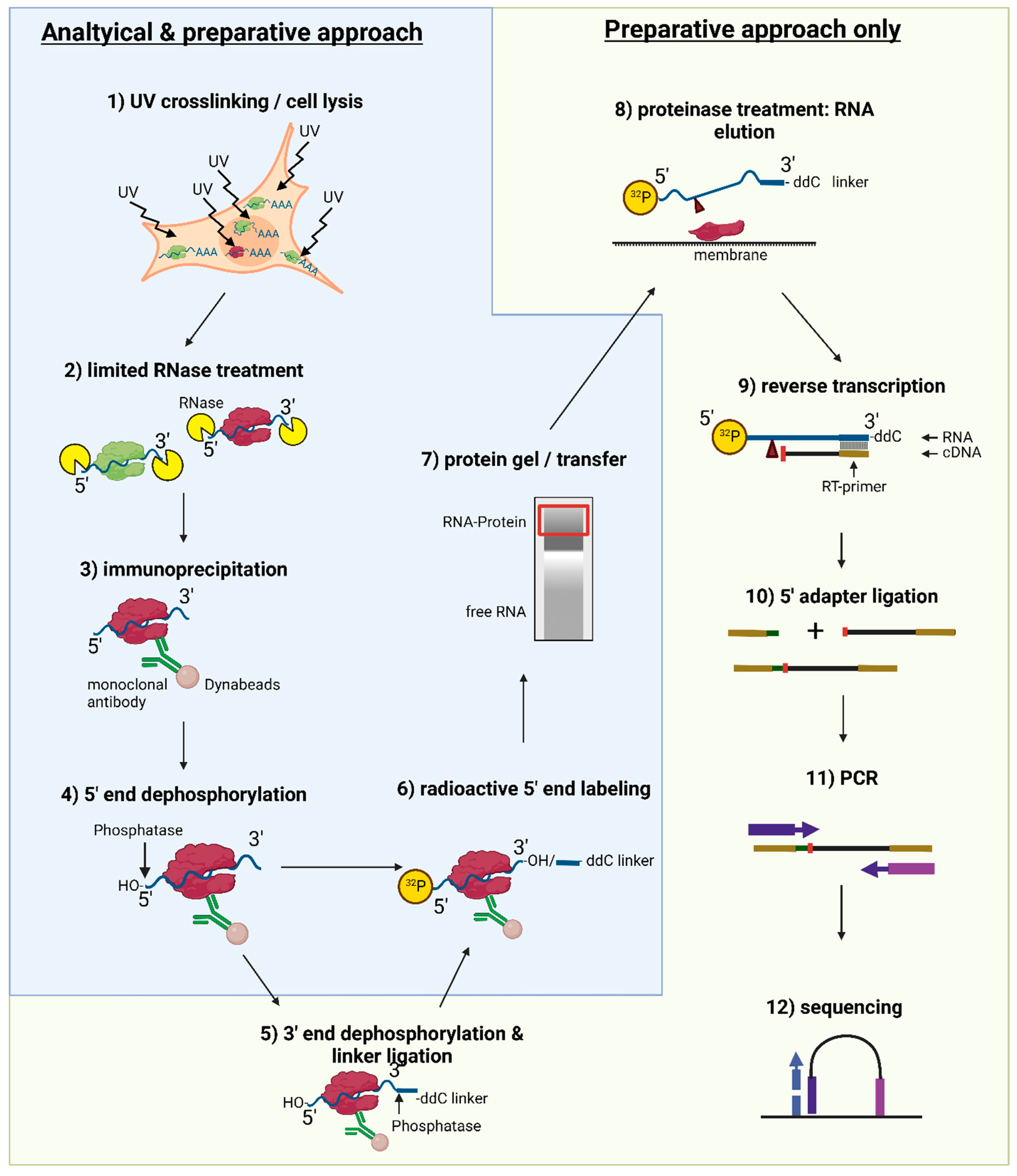
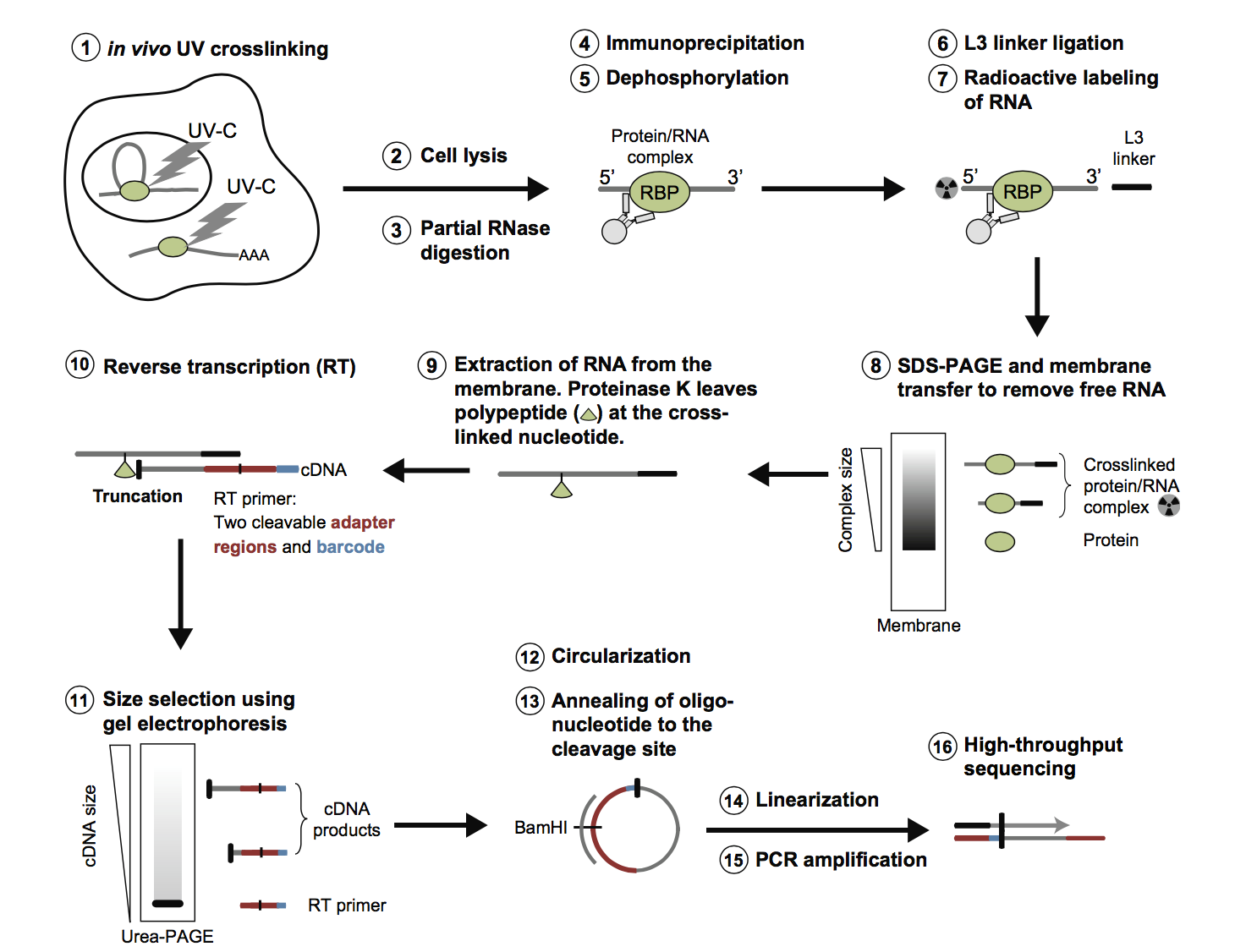
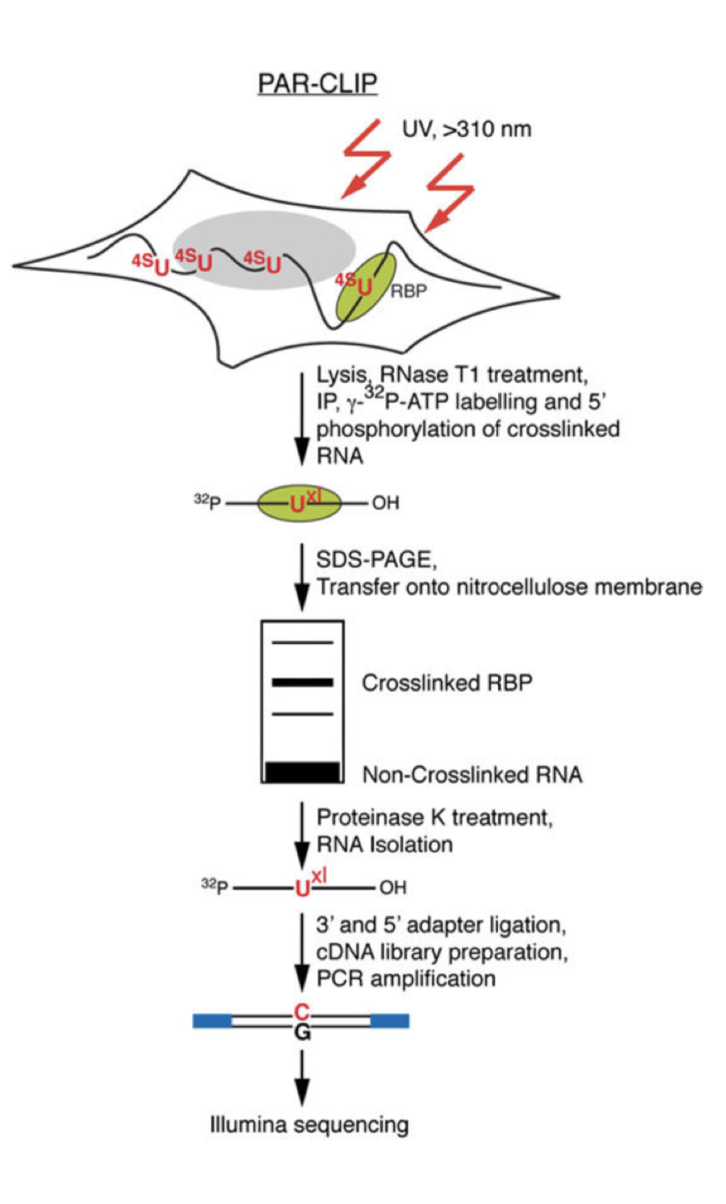
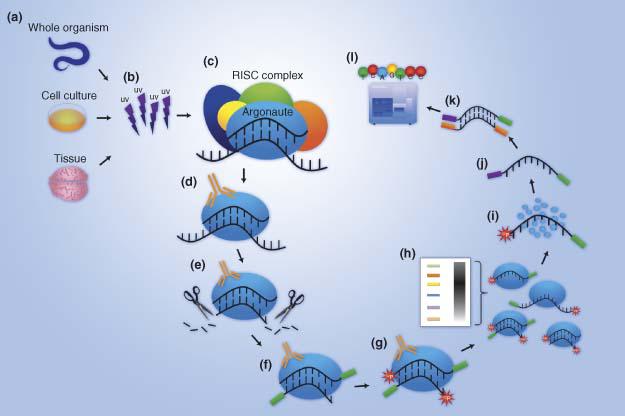
MPs | Free Full-Text | Studying miRNA–mRNA Interactions: An Optimized CLIP-Protocol for Endogenous Ago2-Protein

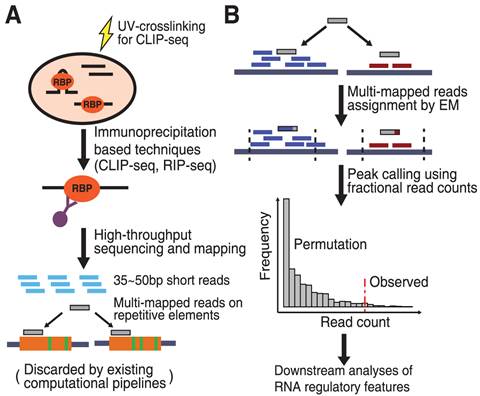
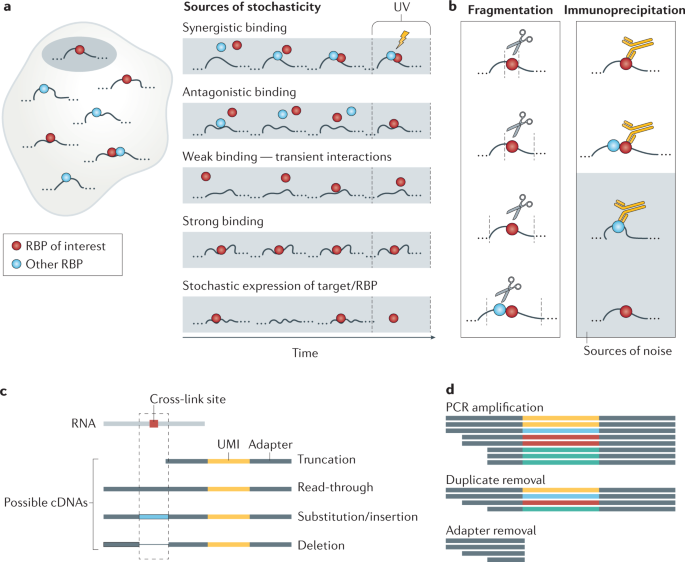
Bioinformatic tools for analysis of CLIP ribonucleoprotein data - De - 2017 - WIREs RNA - Wiley Online Library

-2.jpg)